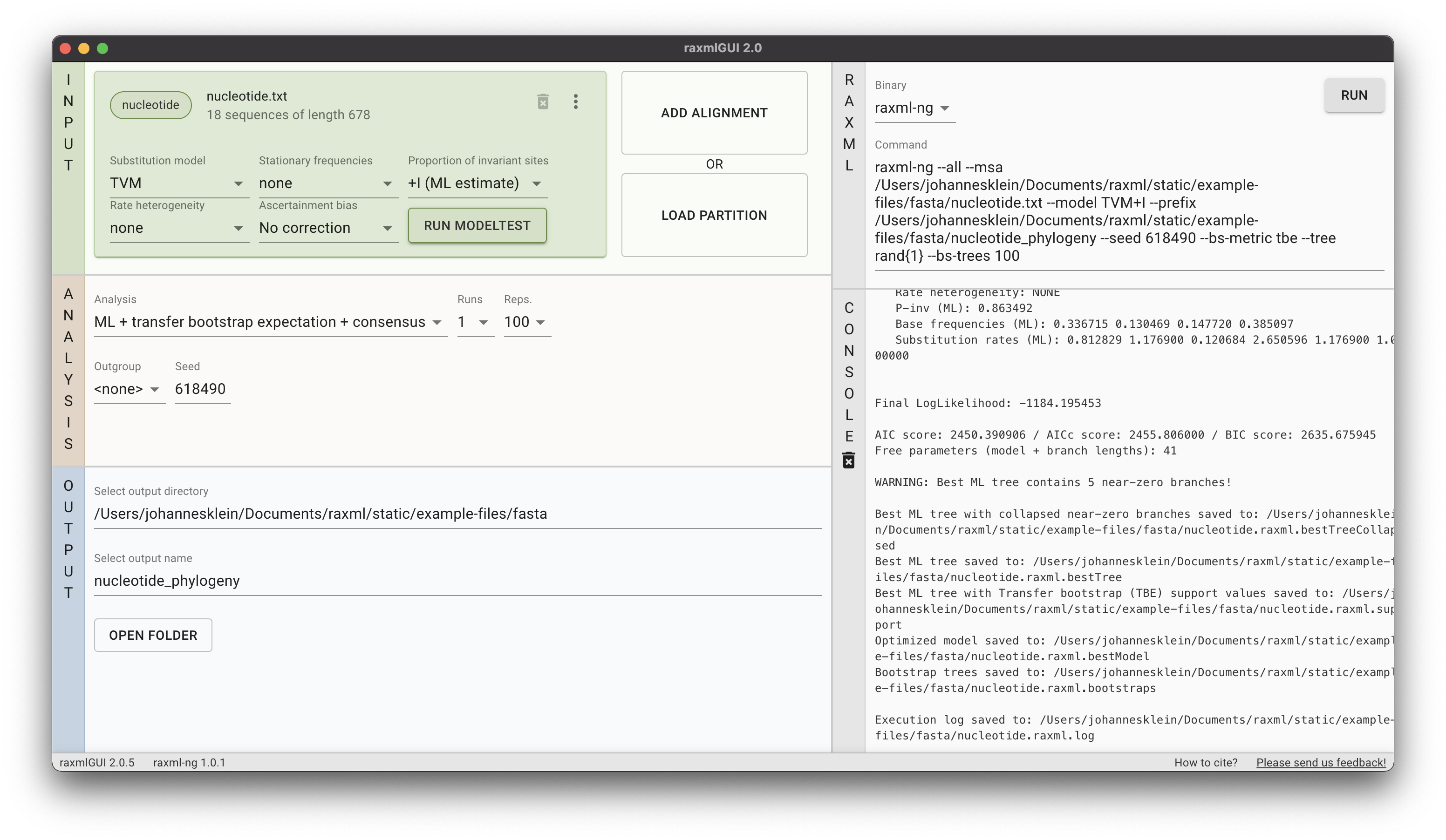
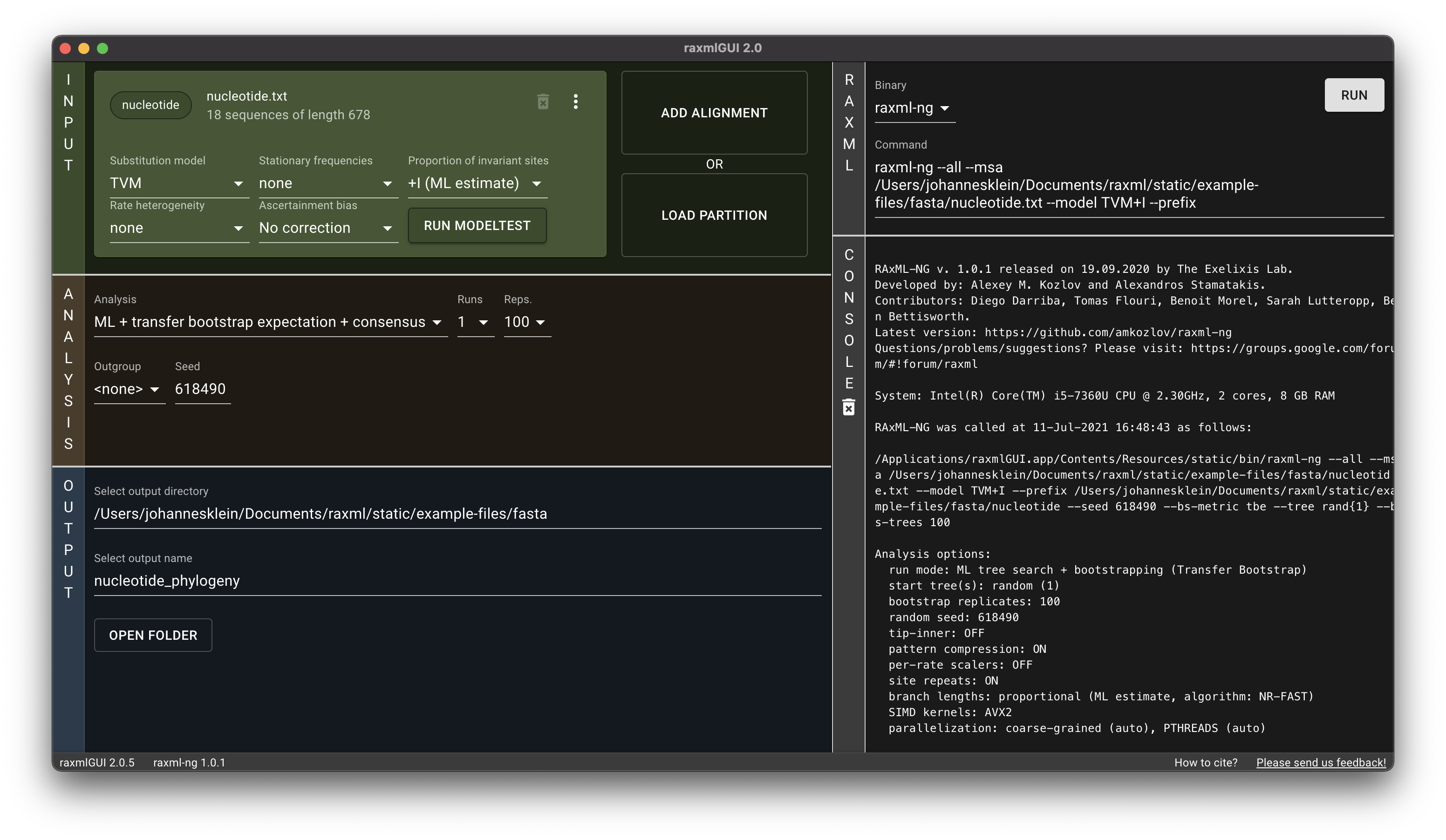
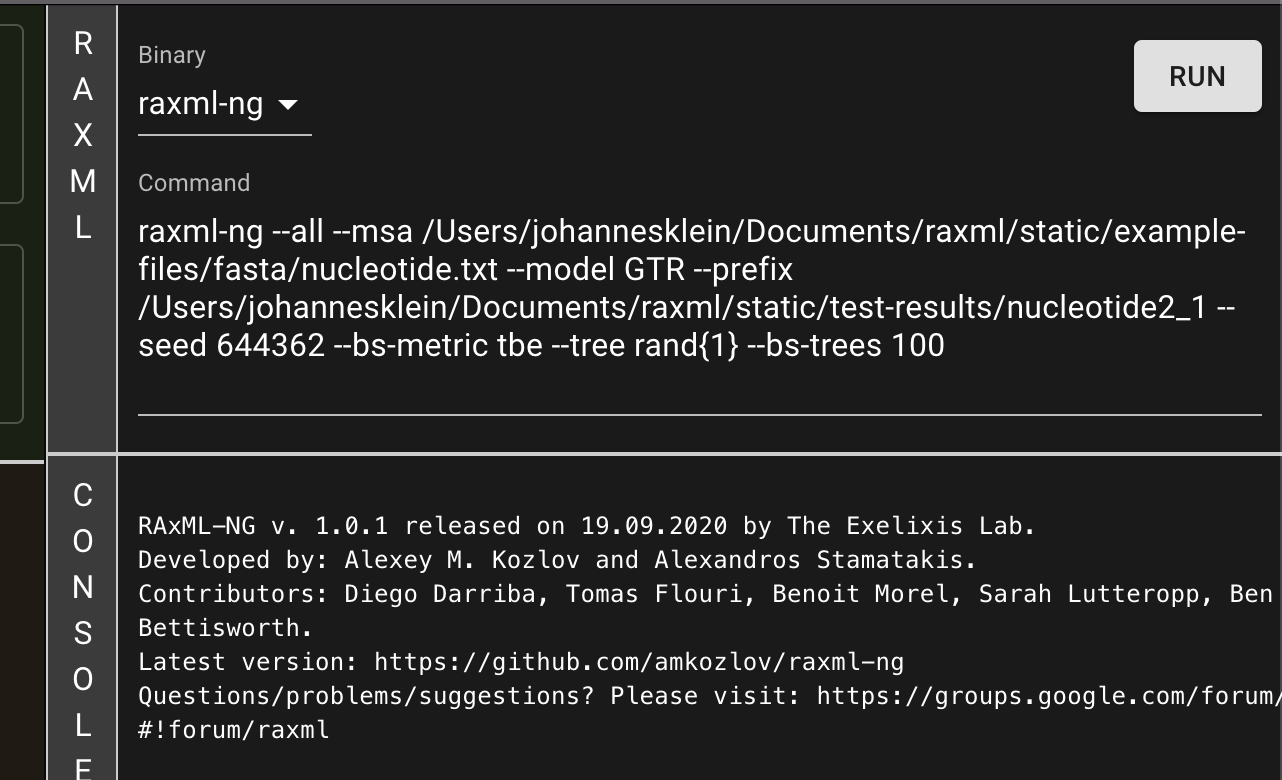
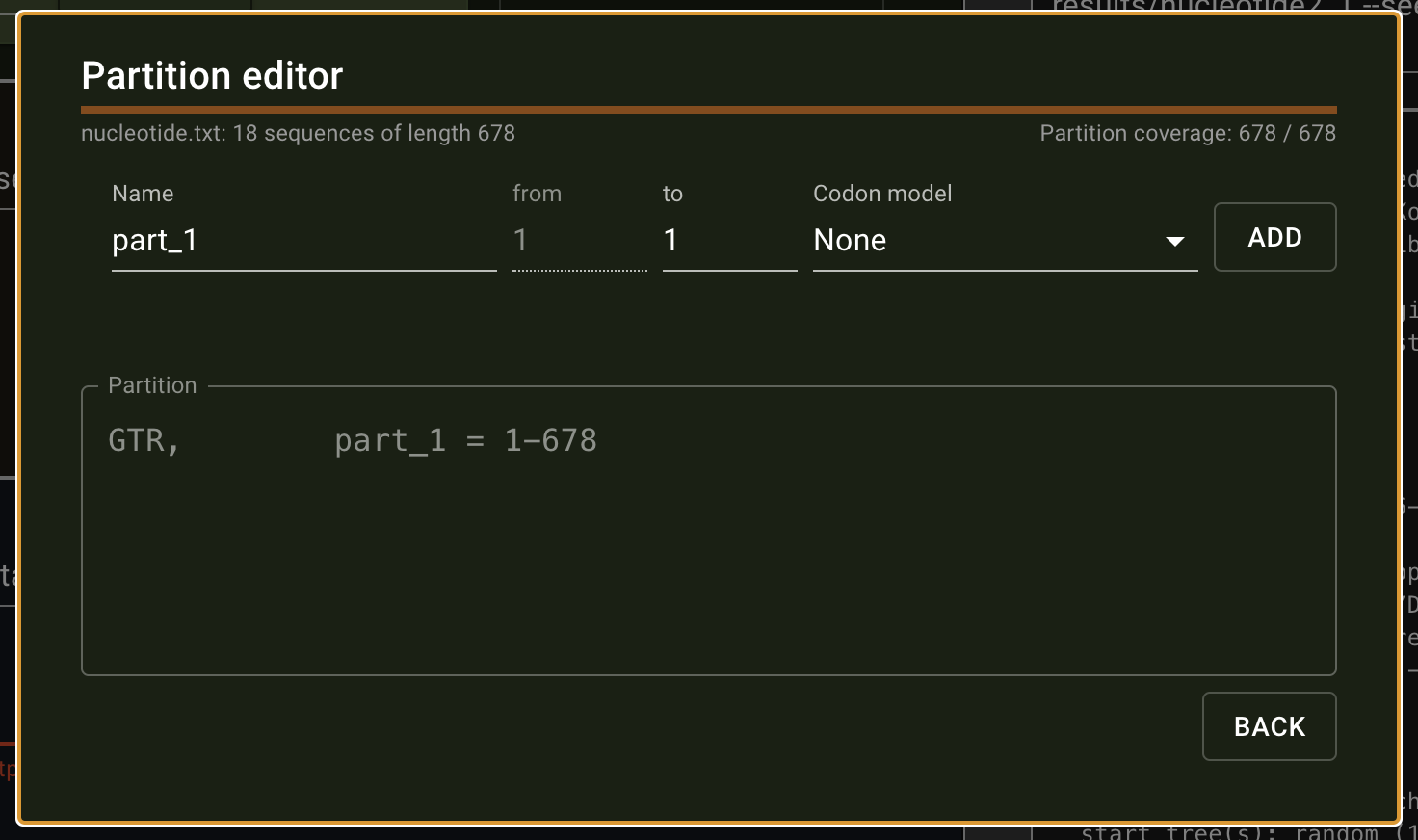
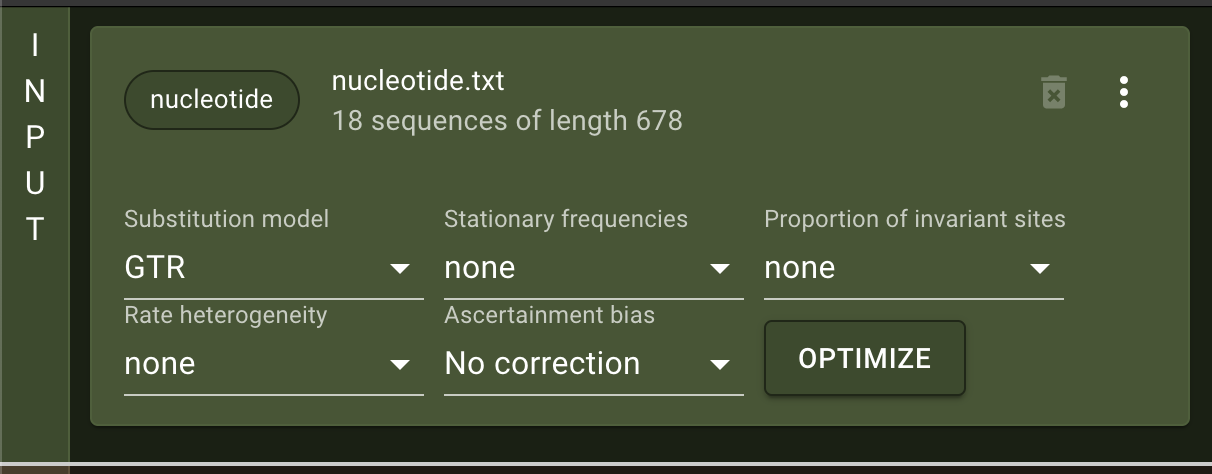
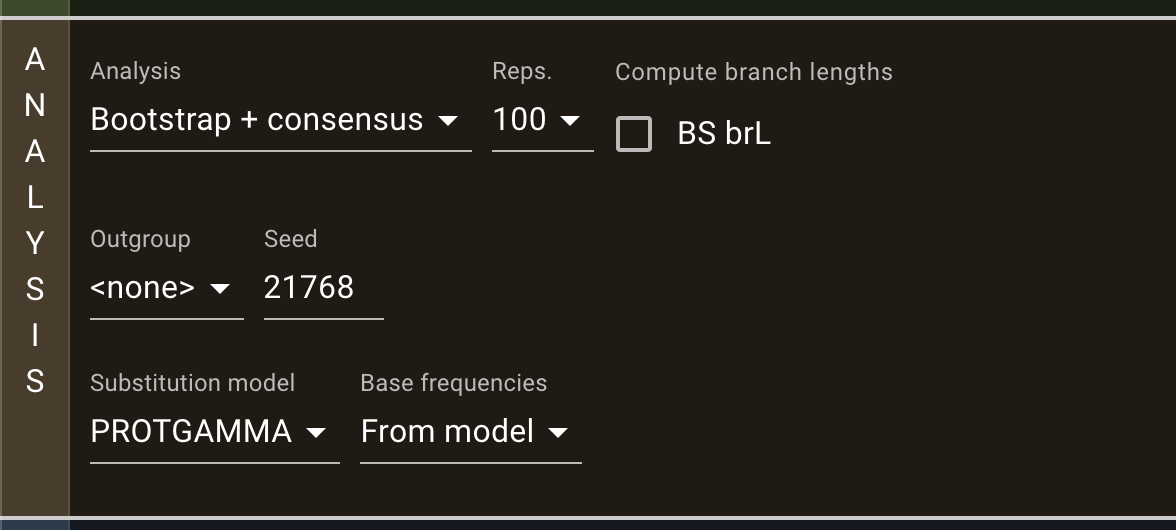
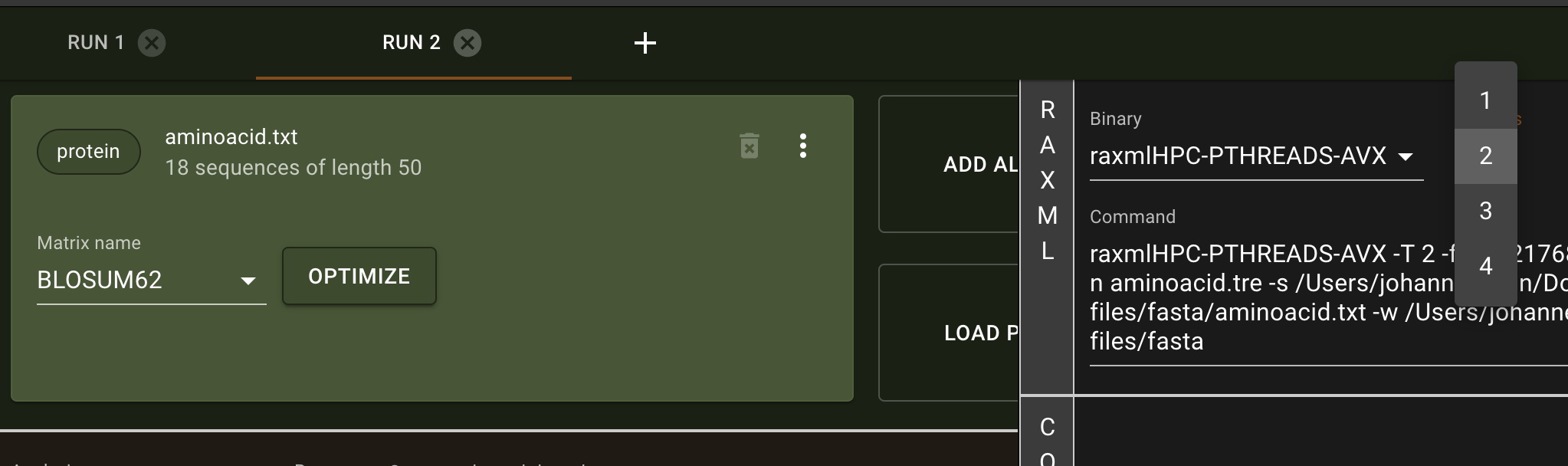
The program presented here is the first release, the most important functions and analyses are implemented and functional. However, we encourage users to send us any feedback they may have. raxmlGUI 2.0 facilitates phylogenetic analyses by coupling an intuitive interface with the unmatched performance of RAxML. Read more in the accompanying paper.
1 Edler, D., J. Klein, A. Antonelli, and D. Silvestro. 2020. raxmlGUI 2.0: A graphical interface and toolkit for phylogenetic analyses using RAxML. Methods in Ecology and Evolution, doi: http://dx.doi.org/10.1111/2041-210X.13512
2 Silvestro, D. and I. Michalak. 2012. raxmlGUI: a graphical front-end for RAxML. Organisms Diversity and Evolution 12:335–337.
3 Stamatakis, A. 2014. Raxml version 8: a tool for phylogenetic analysis andpost-analysis of large phylogenies. Bioinformatics 30:1312–1313.
4 Kozlov, A. M., D. Darriba, T. Flouri, B. Morel, and A. Stamatakis. 2019. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455.
5 Darriba, D., D. Posada, A. M. Kozlov, A. Stamatakis, B. Morel, T. Flouri. 2020. ModelTest-NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models. Molecular Biology and Evolution 37:291–294.